Analyze

Automation
Automation link with SigmaPlot 2000, Excel 9, Origin 6 or 8.1 or MATLAB >v.5.3. Program usage must be enabled in the ini file under the 'Script' tab. Linking to earlier versions possible if supported by those versions. If application icons are present in the upper right of the jClamp main page when in analysis mode then the programs are available for use. When active, drag icon in upper right of main jClamp window onto a graph and release -- in Analysis Window, IV-Plot Window or CC Data Plot window -- and data is automatically transferred to the application and a plot is generated. SigmaPlot templates (*.JNT) can be stored in the "jClamp32/sigmaplot" directory for use of user defined graphs. Upon dragging and releasing the SigmaPlot icon, a list of template files is displayed and can be chosen. Application files should be saved under new names before exiting jClamp or inactivating the application if data loss is to be avoided. MATLAB commands can be run from analysis scripts. SigmaPlot transforms can be run from analysis scripts. Origin LabTalk command can be run from analysis scripts. All applications communicate with jClamp via Active X (Ole automation) except the older Origin program which only supports slow, legacy DDE. Because of this, the full path to the origin executable, e.g., "c:\microcal\origin60.exe", needs to be entered into the jClamp ini file in the 'Script' tab section. Otherwise the application will not be recognized.
Automation links remain in effect when exiting Analysis mode. This allows quick reentry to analysis mode rather than checking for connections each time. No need to double clik on the Sigmaplot, Matlab, Excel or Origin icons in Analysis mode. If programs are available (but need to be set to check for [in in file]) then icons will display and drag/drop icons to transfer data will be immediately available.
In order to use SigmaPlot jClamp templates, available in jClamp SigmaPlot directory, you must load them into the template.jnt files according to Sigmaplot instructions. The default template.jnt is located in the SigmaPlot program directory (see below).You can add templates for access in jClamp by creating your own in SigmaPlot and putting the names of the template in the jclamp/sigmaplot directory (e.g., make a file name "my_new_template.jnt" in the directory -- it can be an empty file, only the file name is necessary and must match the template name "my_new_template" in the sigmaplot template.jnt file. Check that the new template is available in SigmaPlot. It is required that you reload SigmaPlot to install new templates that are added to template.jnt.
Sigmaplot transfers will now increment notebook numbers so that now saving with new names is not required between jClamp->SigmaPlot transfers. jClamp data file names are inserted in notebook sections.
All transfers of data to other program via automation will leave the automation program files open after jClamp closes. You can continue work on the data in those programs (SigmaPlot, Matlab, Excel or Origin ) and save what you like. Matlab will ask before closing, Xcel will ask to save data before closing. SigmaPlot will not close.
Analysis Window
A large window displays the data file traces and permits certain types of data manipulation. Descriptions of the button functions are provided in the lower main window panel by moving the mouse over the buttons. Data is loaded with the file open button in the task section. Other options include: plot printing; I-V analysis; zoom viewing; Cell Censor data viewing; toggling of leakage subtraction; exporting files into abf format, Matlab format or ASCII format; clipboard capture of the displayed traces data or graphic plot; file math capabilities; and episode math capabilities;.
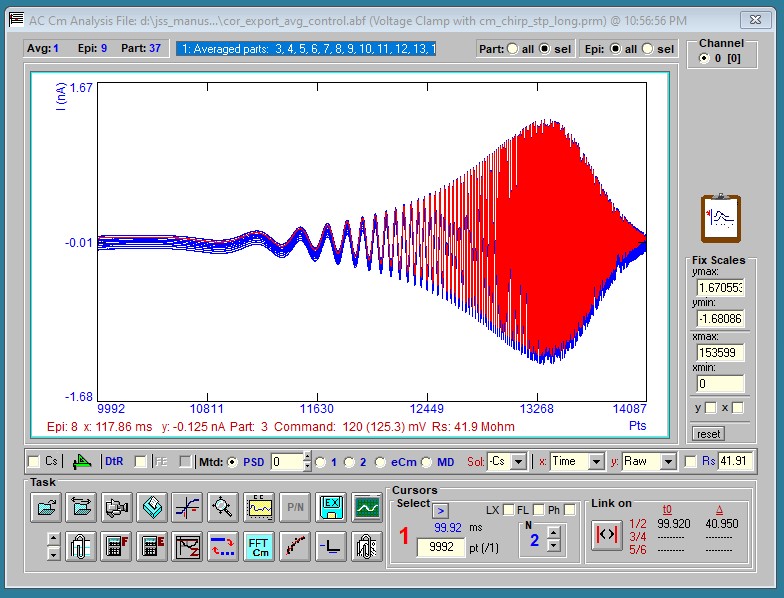
Various plotting manipulations are possible with the mouse. Particular episodes and/or partitions can be plotted, or all data can be plotted. Vertical scaling of traces is accomplished by placing the mouse to the left of the y-scale where it changes to a magnifying glass image; press the left mouse button down and drag it vertically until the desired selection is made (indicated by a colored bar along the y-scale). Upon releasing the button, the traces will be magnified. Horizontal scaling is similarly done beneath the lower x-scale. Double clicking on the plot or pressing the reset button will restore full trace scaling, if Fixed Scales are not enabled. Fixed X and/or Y scaling is selectable By pressing the left mouse button down within the graph area, which displays a open hand cursor, and dragging, the traces can be shifted around. A small plot in the lower right shows the horizontal position during dragging. Movements of the mouse over traces highlights the selected trace, and displays information at the bottom of the plot. Vertical movements of the mouse to the right of the right y-scale displays a horizontal cursor over the plot with a text box indicating magnitude. Y-scale values are changeable by clicking on them or pressing <shift><Y> -- current scale loops through nA, uA, mA, and pA . Default scales are set in the ini file. X-scale values can be in points (pts) or time (ms). Clicking on the x-scale label or pressing <shift><X> toggles between the two.
Plots can have axes or a scale bar. The scale bar will indicate magnitude for the plotted data. The scale bar can be dragged to any location in the plot window. Select the corner of the scale and a finger pointer appears; drag very slowly with the mouse down to a new location. In the scale bar mode, voltage protocol is displayed as well. In addition, if the digital partition output is enabled in the Command window, then digital output is plotted. In either plot mode, a colored marker (up to 3) can be placed on the graph by clicking the right mouse button over the data plot. Left clicking on them will erase them.
The click clip check box if enabled will display a magnifying glass cursor which can be used to capture x,y points from the plot to the clipboard. Points are appended to the clipboard with each left click of the mouse. You can paste the data into any spreadsheet. Unchecking the check box clears the clipboard.
See info panel at bottom of main jClamp window for button info when mouse moves over each button. They are enabled after a data plot is created.
If a ramp command stimulus was employed to collect any part of the data, a panel is displayed to allow i-v plot generation
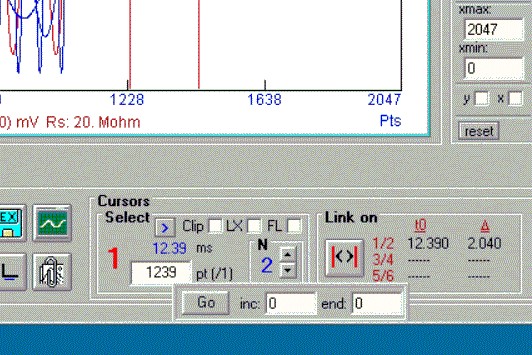
Added auto cursor increment with iv analysis. Selected cursor is incremented and auto analysis in iv window (fit fitting) can be done.
CC Data Plot Window button - use to view saved Cell Censor data (*.cc), capacitance tracking data (*.cmt) , and capacitance data (*.dcm) obtained from protocol files (*.prm) that incorporate two-sine voltage stimuli.
 Two data files can be displayed simultaneously by turning on storage. The previously loaded data plot is retained in the color green, and its filename is displayed in green in the Analysis window.
Two data files can be displayed simultaneously by turning on storage. The previously loaded data plot is retained in the color green, and its filename is displayed in green in the Analysis window.
Cm Analysis in the Data Plot Window
It is now possible to derive Cm, Rs, Rm, and Im from the single, dual or eCm method abf files in the Analysis window. Most analysis features that are used for currents (exponential fitting, etc) are then available to use on the extracted data. When a file that has employed 1 or 2 sine voltage stimulation is viewed in the Open data file window (see that help section), you can enable a checkbox to load the embedded calibration file (or if unchecked, use one already loaded or load a new one for analysis from the Analysis window). A calibration file opened while in Analysis mode will only be used for analysis; the data collection file remains the same, and can be changed only when Analysis mode it exited. Upon plotting the abf file in the data analysis window, a panel is displayed at the lower right that allows choosing which data to plot -- Raw, Cm, Rs, Rm, Im or Vm. You may also choose whether to analyze with single sine, dual sine or the new eCm Method, and correct for series resistance effects on voltage. The green scale icon can be used to load calibration files, and view current settings (in the jClamp info panel at the bottom of the full screen). You may also choose to detrend data or not. You have to read the detrending note if you superimpose sinusoidal stimuli on ramps or other fast time varying commands. See the Technical Note in the Technical Note subdirectory under the jClamp directory for information on detrending. Detrending now corrects without adding any noise to the data.
If you set the angle to measure single sine in CC, it is available here in the PSD box. PSD must be selected to use the angle to correct the magnitude of current at that angle and 90 degrees away, which can be viewed when accessed under the X, Y drop down boxes. Also Real and Imaginary components of the raw current can be viewed.
If you used the Cs feature in CC prior to collecting sinusoidal protocol data, you will be able to correct for the electrode stray capacitance by checking the Cs box (which will only be enabled if you performed Cs correction). The corrections are saved with the data file.
In the Cm analysis pane, the dropdown box “f1/f2” (selection 1,2, or 3) permits the extraction of Cm based solely on ω1, ω2 or the average of the two, respectively. While using a math model produces equivalent results, real world conditions (e.g., different noise levels at each frequency, changes in stray capacitance, nonlinear distortion) may produce different results. Using 1 is probably best. Averaging was internally set for jClamp versions 12.0 – 12.5.
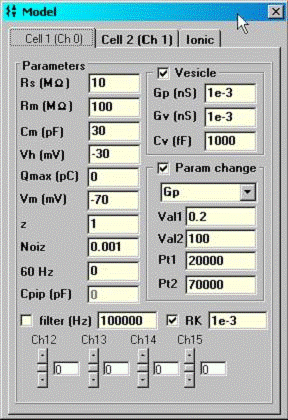
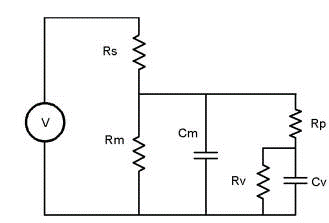
Fusion event analysis is also possible. The following model is used. Basically, prior to vesicle fusion Rp is consider to be infinite and the admittance of the cell prior to fusion is assumed to remain constant during the fusion event. Thus Rs and cell’s admittance is mathematically removed from the circuit leaving only the fusion vesicle for analysis (the manuscript detailing the procedure is in review and will be appended when published). Below a math model simulation is analyzed.
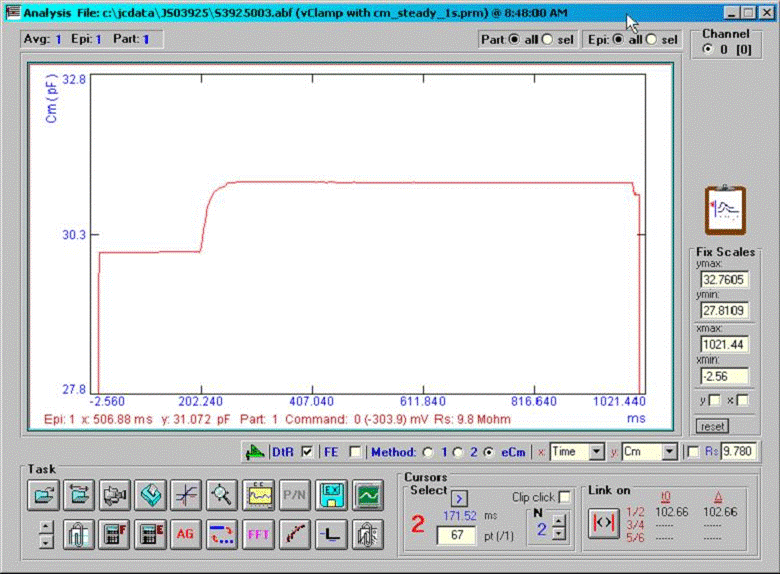
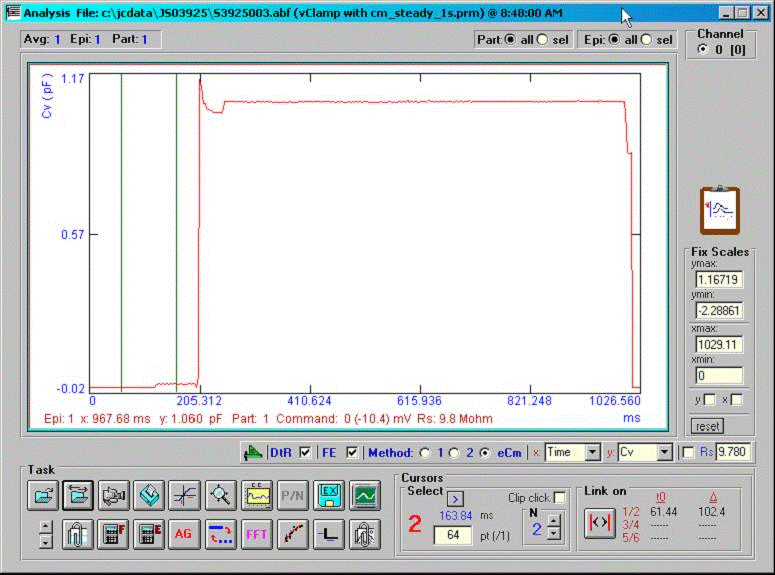
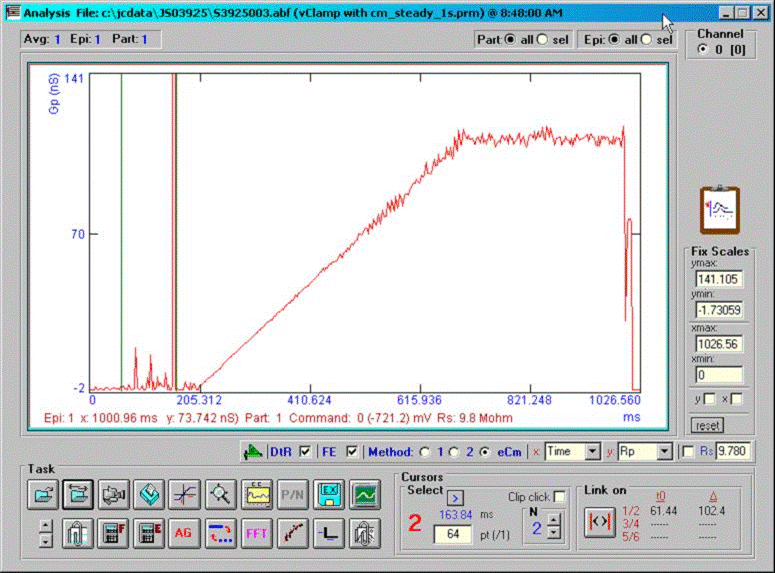
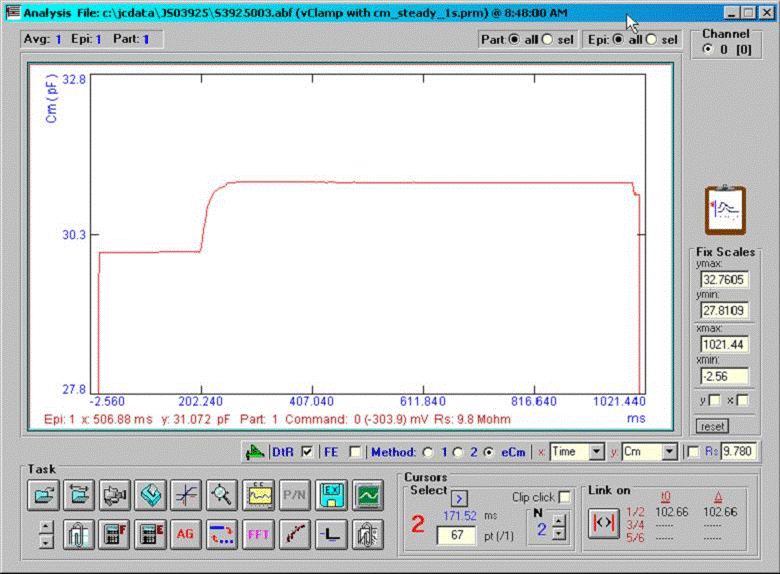
A command protocol which measures Cm during a 1 second interval (10000 points at 10 us clock, 2.56 Cm time resolution ) with the eCm method will be run with the model setup below. With this set up, Gp will initially be 1pS (1000GΩ) at the start of recording; after 20000 points of the protocol have elapsed, Gp will change to 200 pS, which initiates the fusion event. Gp will be ramped linearly between point 20000 and 70000 up to 100 nS, then will remain constant. Below are screen shots of the analysis. If we look at the simple extracted Cm without fusion event analysis, the capacitance changes quickly over the course of 100ms. This time course is not that of the fusion process. If we check the fusion event (FE) checkbox and place single or linked cursors at the section before fusion, we see that in fact the capacitance jumps to final values instantly ( the wiggles are due to the ODE solver). If we now change the plot to the fusion pore conductance (Rp in the dropdown box), and switch to a plot of conductance, Gp (<shift><Y>) we see that the time course of fusion pore expansion is about 500 ms, and the conductance corresponds to the model settings.